|
3/31/2023 0 Comments Splice site mutation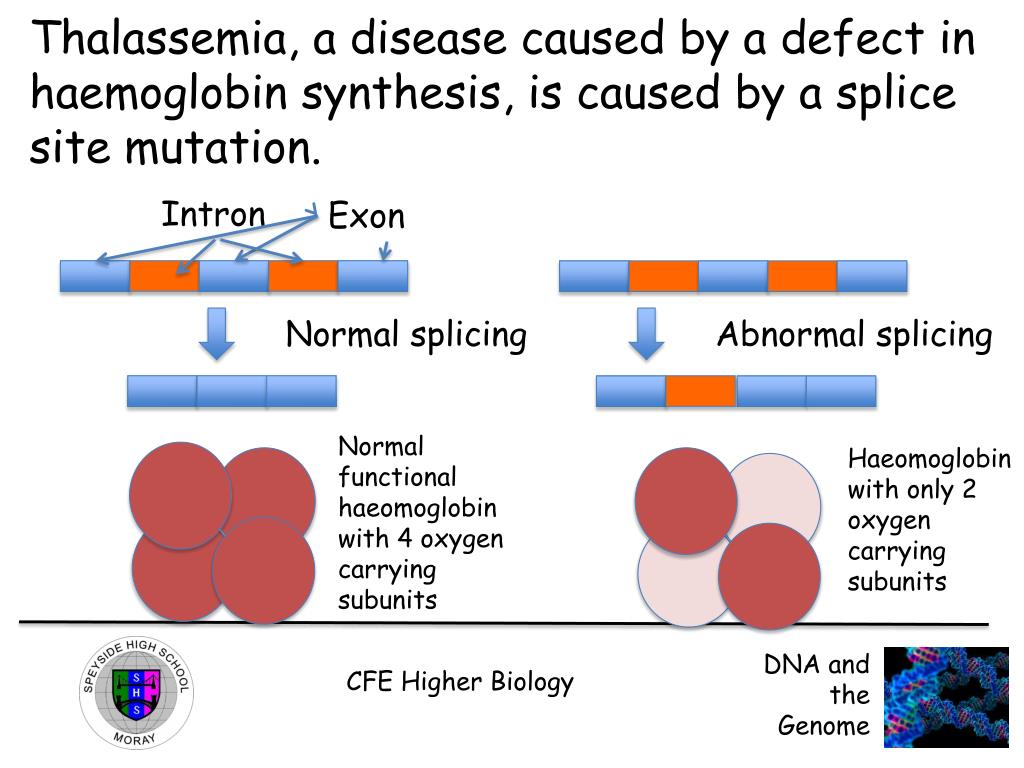
MiSplice searches the BAM file to identify any alternative splice junctions near the mutation of interest, while filtering out known splice junctions and calculating the number of alternative junction-supporting reads for case and control samples. First, the user inputs the locations of RNA-seq BAM files along with a mutation file. (B) The MiSplice workflow consists of three steps: alternative junction discovery, filtering, and manual review. Splice-in is defined as mutations contained within the newly created exons, and splice-out is when the mutation is present in the newly created intron. (A) Examples of splice-site-creating mutations for different conventionally annotated mutation types. RNA mutations of clinical relevance splicing.Ĭopyright © 2018 The Authors. Our work highlights the importance of integrating DNA and RNA data for understanding the functional and the clinical implications of mutations in human diseases. Further, high expression of PD-1 and PD-L1 was observed in tumors with SCMs, suggesting candidates for immune blockade therapy. Notably, we found that neoantigens induced by SCMs are likely several folds more immunogenic compared to missense mutations, exemplified by the recurrent GATA3 SCM. Mutations in 11 genes, including PARP1, BRCA1, and BAP1, were experimentally validated for splice-site-creating function. TP53 and GATA3 have 26 and 18 SCMs, respectively, and ATRX has 5 from lower-grade gliomas. We report 1,964 originally mis-annotated mutations having clear evidence of creating alternative splice junctions. Here, we applied a bioinformatic tool, MiSplice, for the large-scale discovery of splice-site-creating mutations (SCMs) across 8,656 TCGA tumors. For the past decade, cancer genomic studies have focused on mutations leading to splice-site disruption, overlooking those having splice-creating potential.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
